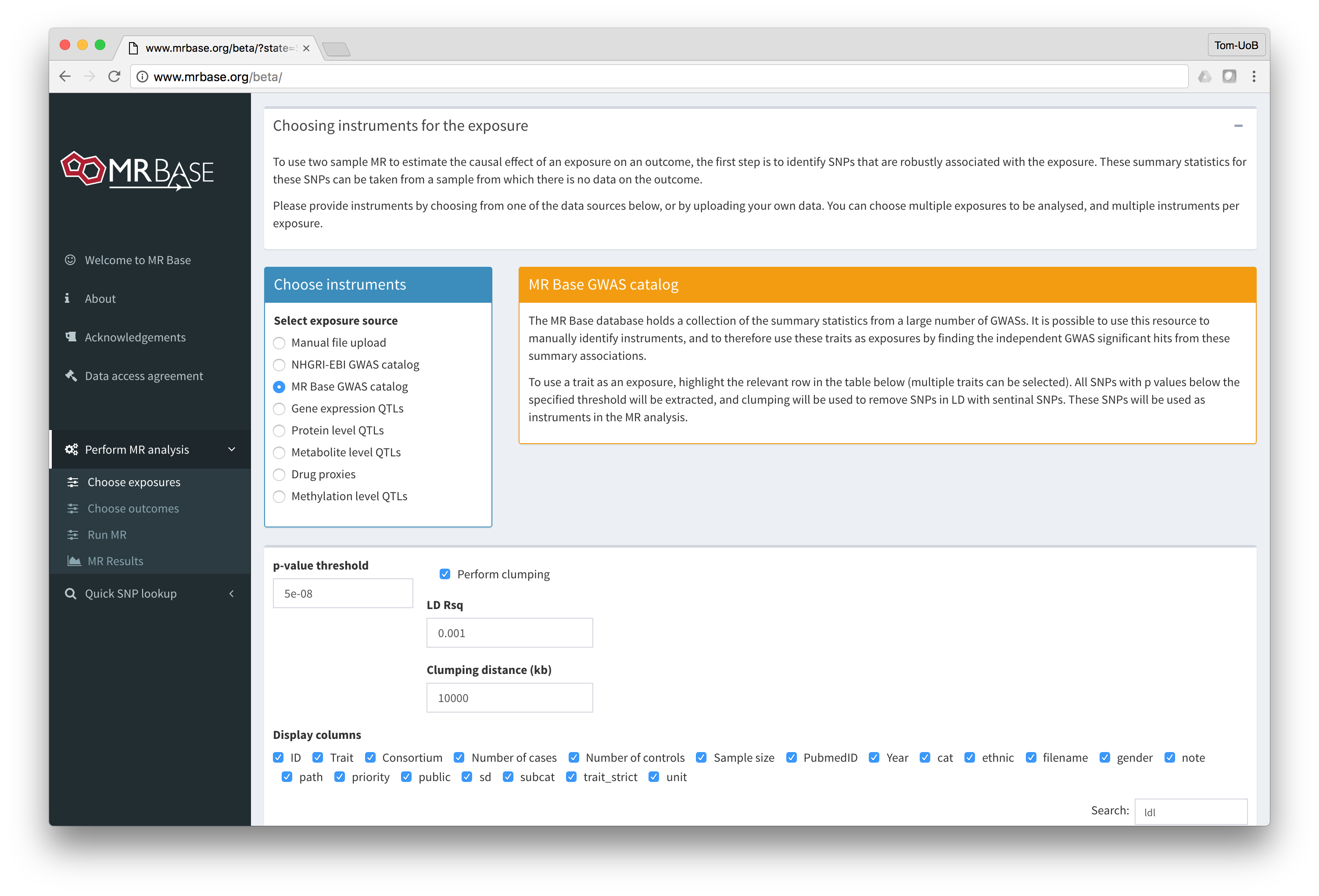
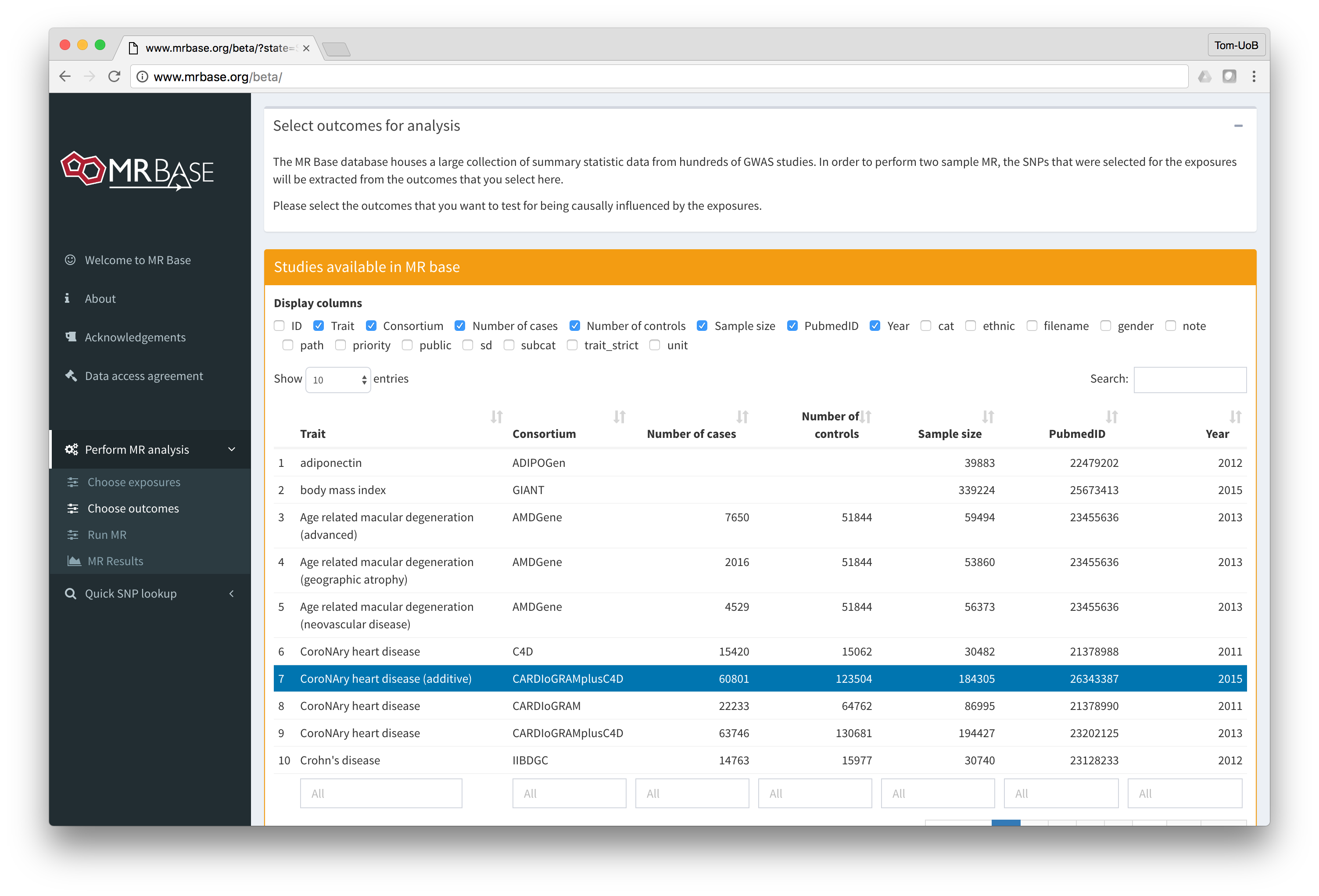
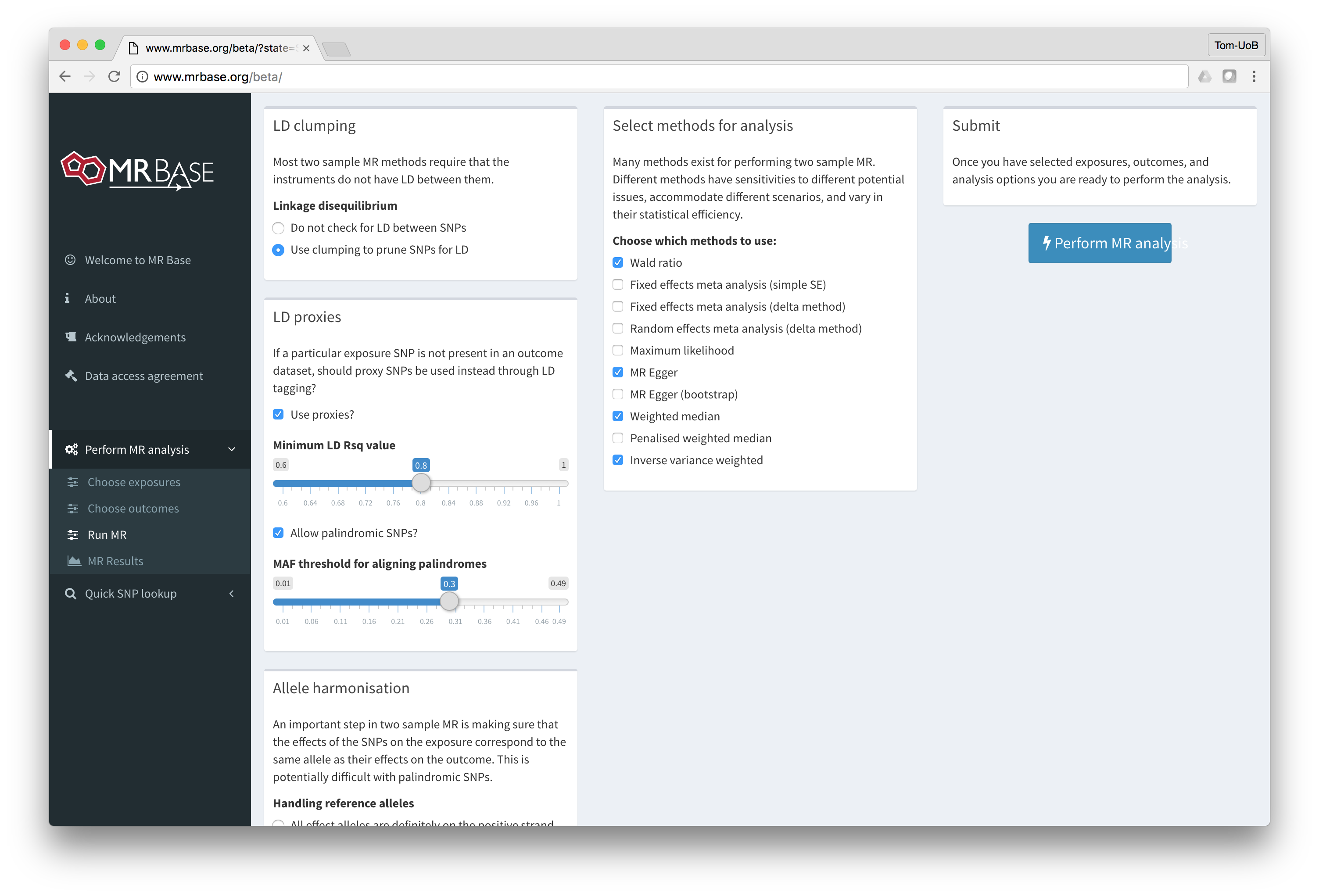
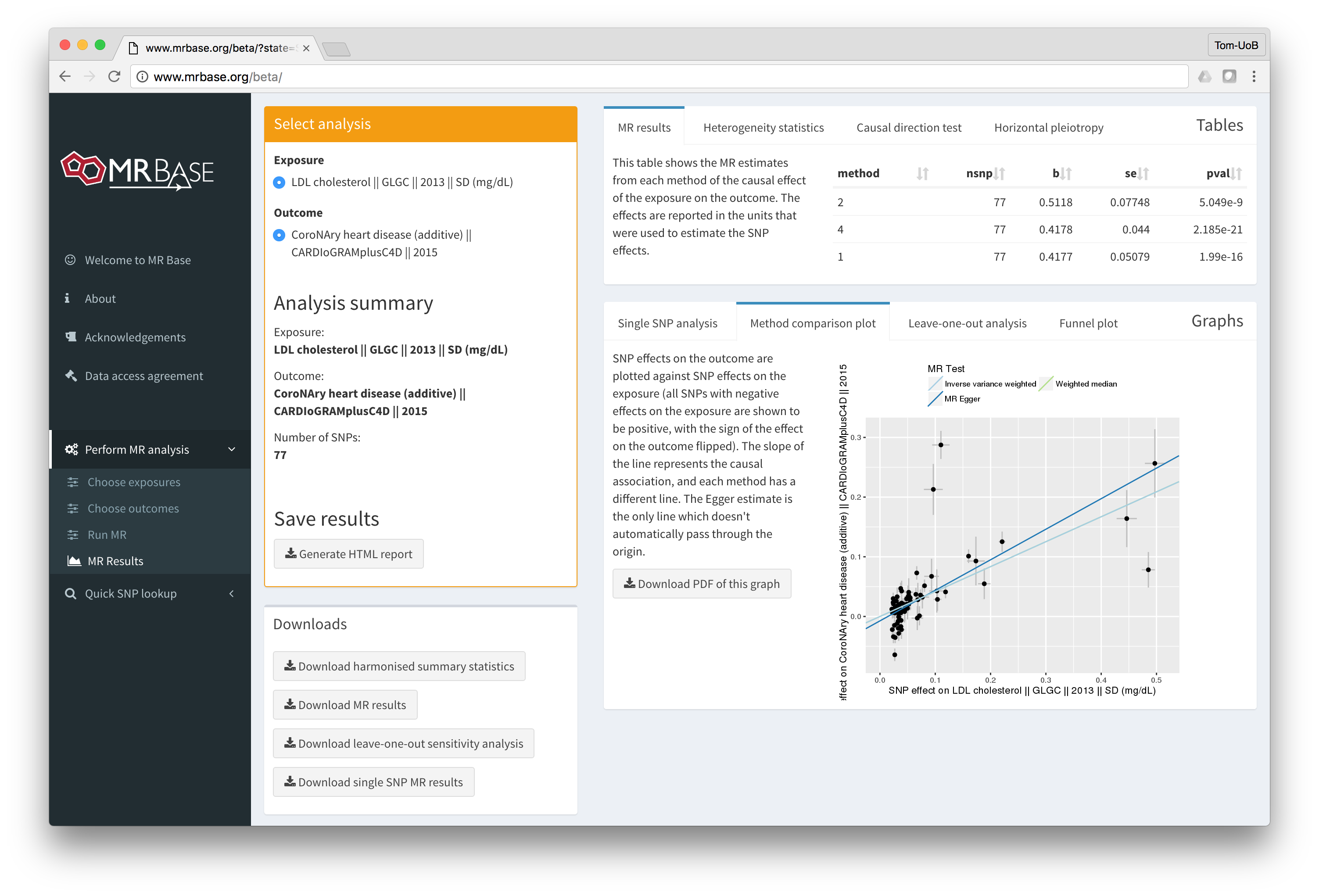
MR-base is a database and analytical platform for Mendelian randomization being developed by the MRC Integrative Epidemiology Unit at the University of Bristol.
You can either use the web application or our TwoSampleMR R package.
Data are also available through the MRC IEU OpenGWAS database.
Launch MR-Base webapp R package OpenGWAS database
Note - by clicking the "Launch MR-Base webapp" button you consent to the use of a cookie which enables us to ensure you have consented to the terms and conditions of data access. Information about how to control or delete cookies can be found at www.aboutcookies.org
The MR-Base paper has now been published in eLife. See the publications page for details.
Our paper reporting Association Between Telomere Length and Risk of Cancer and Non-Neoplastic Diseases has been published in Jama Oncology. See the publications page to access supporting data.
Gibran Hemani, Jie Zheng, Kaitlin H Wade, Charles Laurin, Benjamin Elsworth, Stephen Burgess, Jack Bowden, Ryan Langdon, Vanessa Tan, James Yarmolinsky, Hashem A. $ The MR-Base platform supports systematic causal inference across the human phenome. eLife 2018. doi: https://doi.org/10.7554/eLife.34408
 2-sample Mendelian Randomisation
2-sample Mendelian Randomisation